Bacterial identification can take hours and sometimes longer, treasured time when diagnosing infections and choosing applicable remedies. There could also be a faster, extra correct course of in accordance with researchers at KAIST. By educating a deep learning algorithm to establish the “fingerprint” spectra of the molecular elements of assorted bacteria, the researchers might classify numerous bacteria in different media with accuracies of as much as 98%.
Their outcomes had been made accessible on-line on Jan. 18 in Biosensors and Bioelectronics, forward of publication in the journal’s April situation.
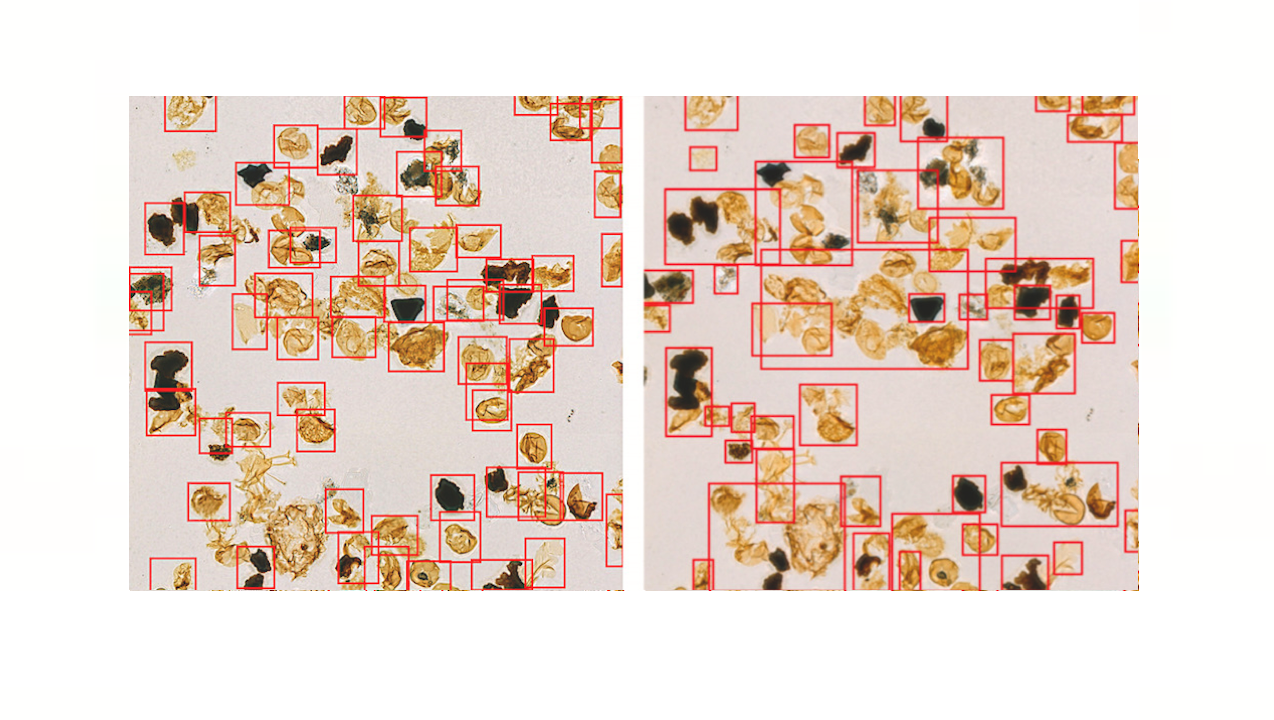
Bacteria-induced sicknesses, these attributable to direct bacterial an infection or by publicity to bacterial toxins, can induce painful signs and even result in demise, so the fast detection of bacteria is essential to forestall the consumption of contaminated meals and to diagnose infections from medical samples, comparable to urine. “By utilizing surface-enhanced Raman spectroscopy (SERS) evaluation boosted with a newly proposed deep learning mannequin, we demonstrated a markedly easy, quick, and efficient path to classify the alerts of two widespread bacteria and their resident media with none separation procedures,” stated Professor Sungho Jo from the School of Computing.
Raman spectroscopy sends gentle by means of a pattern to see the way it scatters. The outcomes reveal structural details about the pattern — the spectral fingerprint — permitting researchers to establish its molecules. The surface-enhanced model locations pattern cells on noble steel nanostructures that assist amplify the pattern’s alerts.
However, it’s difficult to acquire constant and clear spectra of bacteria because of quite a few overlapping peak sources, comparable to proteins in cell partitions. “Moreover, robust alerts of surrounding media are additionally enhanced to overwhelm goal alerts, requiring time-consuming and tedious bacterial separation steps,” stated Professor Yeon Sik Jung from the Department of Materials Science and Engineering.
To parse by means of the noisy alerts, the researchers applied a man-made intelligence technique referred to as deep learning that may hierarchically extract sure options of the spectral data to categorise information. They particularly designed their mannequin, named the dual-branch wide-kernel community (DualWKNet), to effectively be taught the correlation between spectral options. Such a capability is crucial for analyzing one-dimensional spectral information, in accordance with Professor Jo.
“Despite having interfering alerts or noise from the media, which make the overall shapes of different bacterial spectra and their residing media alerts look related, excessive classification accuracies of bacterial varieties and their media had been achieved,” Professor Jo stated, explaining that DualWKNet allowed the workforce to establish key peaks in every class that had been nearly indiscernible in particular person spectra, enhancing the classification accuracies. “Ultimately, with using DualWKNet changing the bacteria and media separation steps, our technique dramatically reduces evaluation time.”
The researchers plan to make use of their platform to check extra bacteria and media varieties, utilizing the knowledge to construct a coaching information library of assorted bacterial varieties in extra media to cut back the gathering and detection occasions for brand new samples.
“We developed a significant common platform for fast bacterial detection with the collaboration between SERS and deep learning,” Professor Jo stated. “We hope to increase using our deep learning-based SERS evaluation platform to detect quite a few varieties of bacteria in extra media which might be essential for meals or medical evaluation, comparable to blood.”
The National R&D Program, by means of a National Research Foundation of Korea grant funded by the Ministry of Science and ICT, supported this analysis.
https://www.sciencedaily.com/releases/2022/03/220304101005.htm